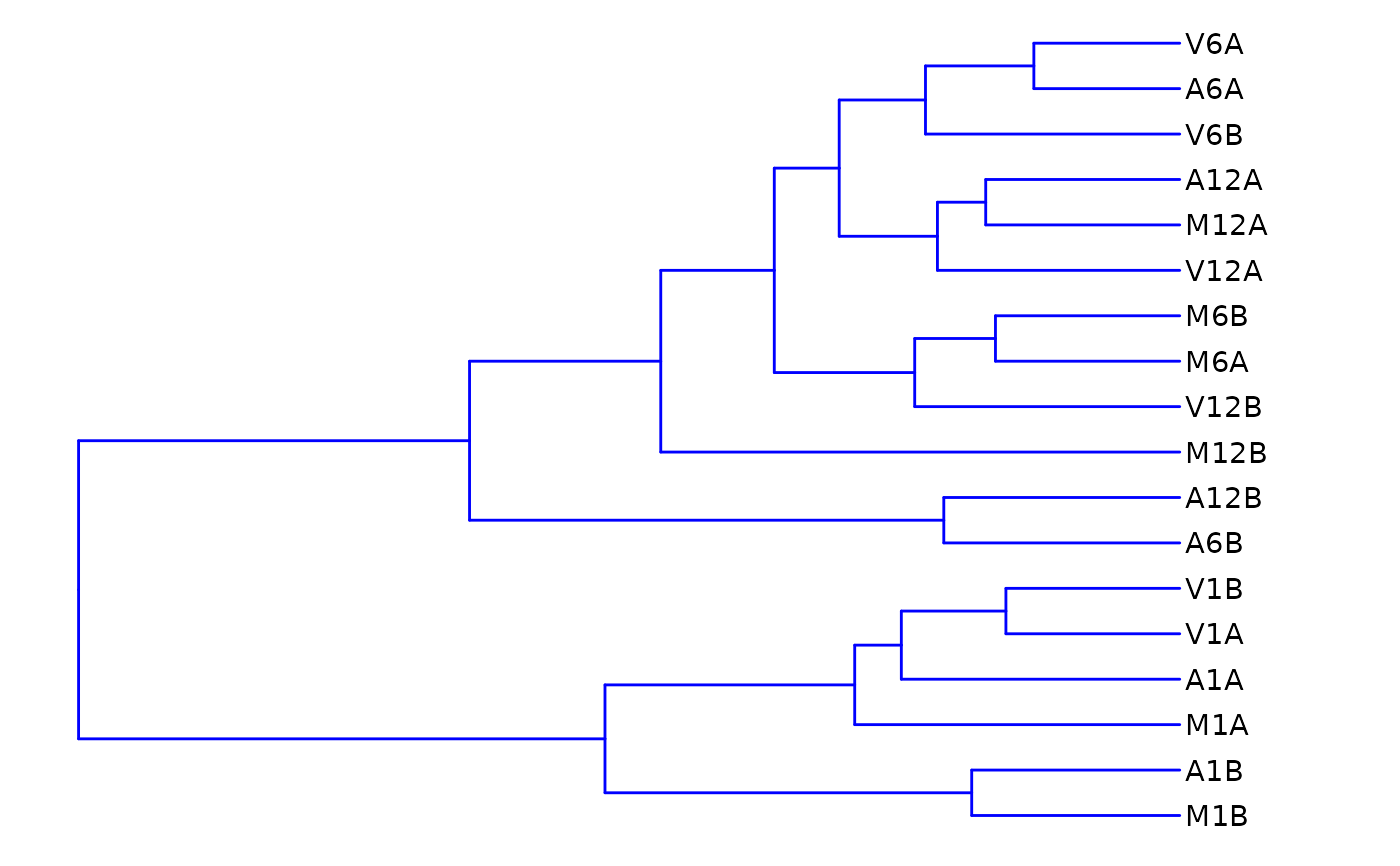
Hierarchical Clustering Dendrogram (hclustplot)
hclustplot.RdThis function computes the sample-wise correlation coefficients
using the stats::cor() function from the transformed expression values.
After transformation to a distance matrix, hierarchical clustering is
performed with the stats::hclust() function, and the result is plotted as
a dendrogram.
hclustplot( exploredds, method = "spearman", plotly = FALSE, savePlot = FALSE, filePlot = NULL )
Arguments
| exploredds | object of class |
|---|---|
| method | a |
| plotly | logical: when |
| savePlot | logical: when |
| filePlot | file name where the plot will be saved. For more information,
please consult the |
Value
returns an object of ggplot or plotly class.
Examples
## Targets file targetspath <- system.file("extdata", "targets.txt", package = "systemPipeR") targets <- read.delim(targetspath, comment = "#") cmp <- systemPipeR::readComp(file = targetspath, format = "matrix", delim = "-") ## Count table file countMatrixPath <- system.file("extdata", "countDFeByg.xls", package = "systemPipeR") countMatrix <- read.delim(countMatrixPath, row.names = 1) ## Plot exploredds <- exploreDDS(countMatrix, targets, cmp = cmp[[1]], preFilter = NULL, transformationMethod = "rlog" ) hclustplot(exploredds, method = "spearman")